Protein backbone angles12/31/2023 
The popular training techniques include neural networks (NN), , support vector machines (SVM), and hidden Markov models (HMM). Įncouraging progress has been made in purely sequence-based backbone torsion angle predictions, where investigators usually divide the backbone conformations into several discrete states based on the phi/psi values and then use various training algorithms to predict the states of variant phi/psi values, , –. In the field of structural bioinformatics, the torsion angle prediction data have found their usefulness in aiding secondary protein structure prediction, , sequence alignment, fold recognition, and protein structure modeling. For example, SHIFTOR and PRIDICTOR developed at Wishart's lab can generate quickly high-resolution predictions of phi and psi values using the chemical shift data and the sequence information. With the development of computing technology, the computer-based algorithms can accelerate the determination of backbone dihedral torsion angles. The experimental procedure of the phi/psi angle determination is usually laborious and time-consuming. Therefore, if the values of the phi (φ) and psi (ψ) angles are known, the geometry of the global protein structures can be readily constructed with the standard bond length. Because of the planarity of the partial-double peptide bond, the torsion angle ω is almost fixed at 180° with rare cis cases of 0°. 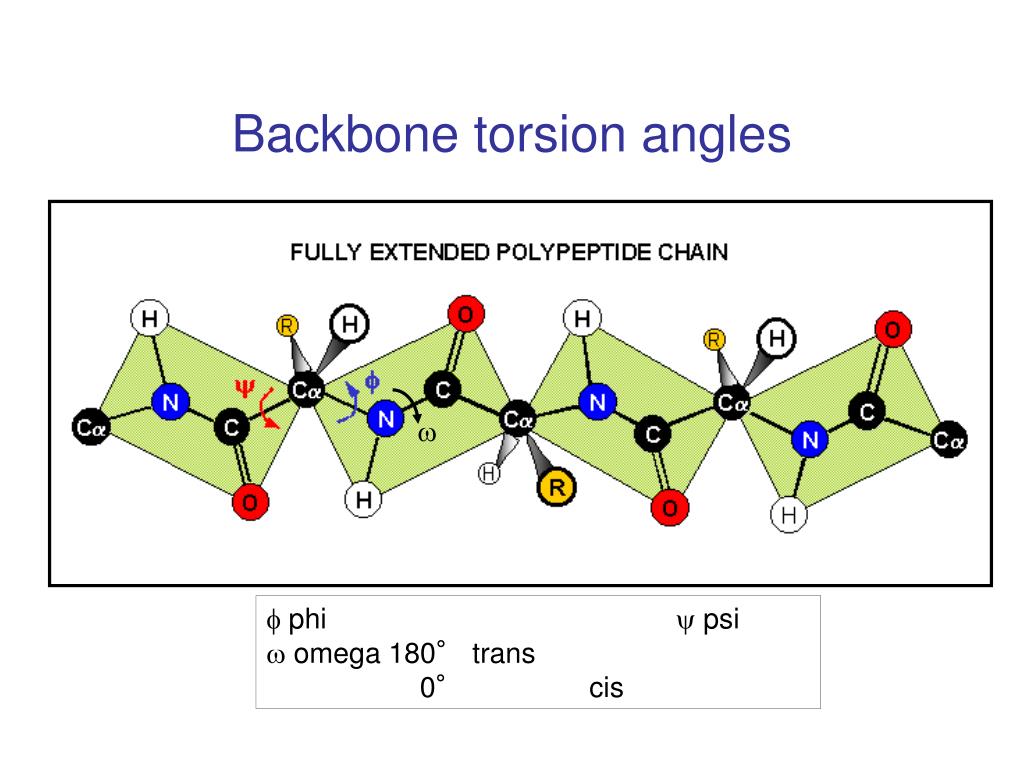
There are three backbone dihedral torsion angles along with the protein peptide chains, which dictate the topology of protein 3D structures, i.e. Thus, although the average accuracy of the ANGLOR prediction is still low, the portion of the accurately predicted dihedral angles may be useful in assisting protein fold recognition and ab initio 3D structure modeling. For some residues (ILE, LEU, PRO and VAL) and especially the residues in helix and buried regions, the MAE of phi angles is much smaller (10–20°) than that in other environments. The prediction is statistically different from a random predictor (or a purely secondary-structure-based predictor) with p-value <1.0×10 −300 (or <1.0×10 −148) by Wilcoxon signed rank test. In a large-scale benchmarking test, the mean absolute error (MAE) of the phi/psi prediction is 28°/46°, which is ∼10% lower than that generated by software in literature. The input features of ANGLOR include sequence profiles, predicted secondary structure and solvent accessibility. 
We developed a composite machine-learning based algorithm, called ANGLOR, to predict real-value protein backbone torsion angles from amino acid sequences.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
